A week or two ago, I wrote a guest post on my friend +Jonathan Eisen's blog where I outlined what I thought were some problems standing in the way of folks utilizing #openaccess #preprint servers. I argued the micro-community building, like we're trying to do at Warburg's Lens, or that the folks at Haldane's Sieve have been doing, was a way forward. In that same post, I put a small survey, from which I learned quite a bit, and I want to share those results with you here.
I've never used google speadsheets before, so I learned some lessons about coding answers (doesn't deal well with age ranges etc.) and I had to do a BIT of recoding. There are lots of ways to look at the data, but I picked two to break out specifically. First some summary statistics though:
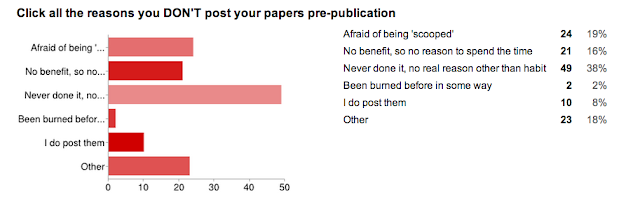
And... the reason we did the whole survey. WHY people don't utilize preprint servers.
I was really surprised by this one. I thought that the 'afraid of being scooped' answer would win the day (this is the case from my anecdotal experience with asking colleagues). I have a good answer to this one, to try to allay fears... but we don't even need it! The number one reason was:
"Pre-print servers? Meh."
And this is easy to change, it is just a culture shift, and one that is, I think in flux (in the right direction!).
You can see all the responses here, if you want, which includes all of the free text responses - where I think the real fodder for change lies. These are hard (impossible) to display, except to copy and paste them all in. But, I can tell you, having read them all, they fall into several categories:
- There is no benefit to me, so why spend the time
*** There is huge benefit, especially to early career folks, as you can show off your work to anyone, with just a link. Further, with the growing popularity of pre-print discussion forums, you can get valuable feedback before submission, hopefully making the process smoother.
- I want to, but my supervisor/collaborator won't let me.
*** Point those folks to these blog posts! Start the discussion.
- I would, if only some big fish in my field did it.
*** The Eisen brothers do it, so can you!
- I'd do it if tenure committees/funding agencies would give me credit.
*** This is a big issue, and I can only hope that the work we do here helps makes that change a reality.
- I'd do it if people could cite my preprints
*** THEY CAN!!! And, google scholar picks up those citations and counts them. Oh, brave new world, with such alt-metrics in it.
- I don't know the rules of the journals that I want to submit to, and I don't want to jeopardize my chances at "real" publication.
*** Just ask Wikipedia - here's the answer
- There's too many choices!
- There's too few choices!
Anyways, please do take a moment to read the responses, they are enlightening. I, for one, have learned that the barriers to getting pre-prints servers more utilized is simply one of outreach - so here I go. If you are in favor of speeding up (and opening up) science, take a minute to engage a scientist in your lab on this issue. It will pay dividends. There are plenty of options for preprint servers now, nicely outlined by +Jonathan Eisen in a post he wrote here, and even more coming on line. I am particularly excited about the bioRxiv coming out of Cold Springs Harbor, and am one of their affiliates in getting going. Until then, I have personally used the physics arXiv for my last five papers as well as a test use of PeerJ which has nice alt-metrics.
Ah, the paradox of choice.
Whatever you choose, post your preprints before you submit. Share your science. Whatever you study, let everyone else see stand on your shoulders so they can see farther, and further the cause.
Monday, June 17, 2013
Saturday, June 15, 2013
Institute for the Future: Health Horizons storification
I was an invited panelist at this year's Institute for the Future: Health Horizons meeting this past week. I had NO IDEA what I was getting myself into when I accepted the invitation, and I have to say, I was skeptical. My experience though, was without question, excellent. Quality organization, quality program, great attendees and a top-notch location. I'll write a proper post to describe the meeting in my own words, but until then, here is a storify version, in tweets. Most are my own, but I used some tweets from others when I was on stage, and also to fill in some spots where I slacked off!
Thanks a million to +Miriam Avery for the invite, and to all my other new friends at the IFTF. I hope to see you all again soon.
Friday, June 7, 2013
Cancer is a sine qua non for life as we know it.
There was a post today on National Geographic talking about a tumor that was found in a fossilized Neandertal's bone. It reminded me that I had written the piece below and hadn't found a home for it yet. The title, which is a big jarring, is:
Cancer is not a disease: there is no cure.
The way we describe things shapes and is shaped by the way that we think of them. Cancer has been described as a disease for as long as we have written record of medicine. It’s name comes from the greek word for crab, because the way it wedges itself into the host tissue is so like a crab wedges itself between rocks: inextricably.
The advent of the microscopic age at the turn of the 20th century brought with it an unprecedented view of the cellular level; anatomy and pathologies came into a new focus and a new science was born: microbiology. As physicians, we were offered a new opportunity to study diseases at the cellular level, to describe the panoply of new patterns that we saw under the microscope much like a team of explorers coming into undiscovered jungle filled with undescribed flora and fauna. We found that the cancers in each organ were not necessarily the same. Indeed, we found a rich diversity of cancers that could be reliably classified and whose prognoses and patterns of progression correlated. This richness of classification gave rise to the opportunity for disease specific treatment trials, and indeed accounted for most of our progress against these individual entities, and for the standard of care for most cancer types even today.
The dawn of the genomic age, first with the human genome project, and then with the cancer genome atlas, promised and delivered another wave of discovery and deeper, more detailed classification. Just as we can now tell how, and when two species of finch, or cave fish, diverged in their evolutionary history, so too can we tell when and how a tumor diverged from its tissue of origin. Early on in this story, we were tantalized by the discovery of specific genomic errors (mutations) that seemed to explain a cancer’s growth, and with the discovery of imatinib, a targeted ‘cure’ for a specific cancer (CML), and cancer seemed to be on its knees, ready to be cured.
The final cure, however, has continued to elude us. As we continue to discover more and more specific mutations, drug companies continue to develop specific drugs to target their action. Each of these drugs seems to work well in a subset of patients, for a time, but never provides the silver bullet that we have been promised, and ultimately fails in almost every case. The problem is that we are stuck in a paradigm where each disease has a cause, and each cause has a remedy. This linear thinking has dominated medicine, and indeed much of science, for most of human history, and has served us well. But continuing to think of cancer as a disease in this paradigm is not going to get us any closer to a cure - we have to shift our thinking and expectations, and embrace the reality that cancer is the result of a non-linear, highly degenerate process, and therefore has no 'cure'.
We have to shift our focus in the study of the cancer genome and stop trying to develop a comprehensive list of errors in the code that cause cancer, but instead learn the guiding principles behind the process - the equations of motion, if you will. Cancer is not a disease to be cured, but a pathologic condition of normal tissue evolving according to the very rules which allowed us to emerge from the primordial ooze. It is an inconvenient sine qua non for existence in our universe, as evolving, living organisms.
This reclassification is not intended to take anything away from those living with cancer, or who have suffered from it. Within any single patient, this pathologic condition has the capacity to cause as much, or more, suffering than does any other disease. And, as oncologists, our calling is to minimize this suffering, and when we can, cure our patient. But until we stop thinking about cancer as a disease that is the product of a linear process, and realize that it is a pathologic condition that is produced by any number of trajectories across an evolutionary landscape; until we stop looking for a single, silver bullet cure for all patients that doesn’t exist, we will continue to waste time that we could be using to understand the evolutionary dynamics of cancer so we can develop strategies to cure each patient.
Some further reading:
Exploiting ecological principles to better understand cancer progression and treatment
arXiv preprint
Basanta and Anderson
Cancer attractors: a systems view of tumors from a gene network dynamics and developmental perspective.
Huang et al. Semin Cell Dev Biol. 2009 Sep;20(7):869-76. doi: 10.1016/j.semcdb.2009.07.003
Oxidants, antioxidants and the current incurability of metastatic cancers.
Jim Watson, Open Biol. 2013 Jan 8;3(1):120144. doi: 10.1098/rsob.120144.
Wednesday, June 5, 2013
NIH Loan Repayment Program
When I was trying to decide what to do with my life after the Navy, it came down to two possibilities: medicine or physics. One, medicine, was wholly new to me but seemed like a good fit for my personality and it was a pretty sure bet, career wise. The other, physics, had been my passion since I was introduced to it in 1992 by Bob Shurtz, my high school AP Physics teacher (and the greatest physics teacher of all time). I had a really hard time picking between the two and ended up choosing medicine for pragmatic reasons, but always hoped to be able to incorporate my scientific inclincations with my new career. I found out pretty late in the game about the MD/PhD programs funded by the NIH through the Medical Scientist Training Program, but applied to a few anyways and was rejected. So, I started medical school as Case Western and was happy.
I distinctly remember one day during MS-1 when I was shadowing a hand surgeon in the OR at the Louis Stoke VA (our teaching VA in Cleveland), and the surgery resident, a PGY-2 in general surgery, described the NIH Loan Repayment plan to me. He was going to take a few years off after his PGY-3 year to pursue a PhD in tissue engineering. The benefit of doing this, he said, was that he could pick his clinical interest first, then pursue a related research area, rather than the MSTP folks who have to pick a scientific discipline before they know what sort of doctor they'll be. This has come into sharper focus for me as my best friend from medical school did his PhD in cardiac stem cells, assuming he would do cardiology, and fell in love with urologic surgery. It isn't so much that his training was wasted, it certainly wasn't, but he lost the momentum that he could have had had his clinical interests been aligned with his doctoral studies. So, in this sense, I'm glad that I wasn't accepted into the MSTP programs, as by going straight through medical school, I found my 'calling' as an oncologist, and then subsequently found how research fits into that - and this is an important distinction.
So, to continue the story, I chose radiation oncology, and now, between PGY-4 and 5, I have been taking time off to do dedicated translational research, and because of the NIH extramural loan repayment program, I am paying off a year of medical school for each year of qualified research! (Considering the opportunity cost of not practicing radiation oncology, I am still a bit 'behind' each year I postpone taking a faculty job, but this dulls the pain significantly.)
While I was disappointed that I didn't get in to the MSTP programs, I truly feel that, at least for me, this path is one that will better suit me for a rewarding career. Obviously, each person is different - had I had a burning passion about a certain kind of research early on, then following that path when I was younger might have been easier. I certainly face a whole slew of personal difficulties as a 37 year old PhD student with 2 young kids that I wouldn't have had I pursued this earlier. On the other side of the coin, with nearly ten years of medical experience, I bring a lot more to the table for my research than I would have after MS-2. At the end of the day, I am happy that the NIH funds both pathways, and we are in dire need of more physicians doing science. Like I said in my TEDMED talk, I truly feel that the physician-scientist plays a unique role in science. Indeed we are specifically trained to connect seemingly disparate pieces of data to build a theory, like a constellation of symptoms to build a diagnosis. As science gets more and more specialised, and knowledge silos get deeper and deeper, I feel this role will grow in importance as well - highlighting the need to keep these programs going.
While I love science and research, I came out of medical school with ~$200,000 in debt. I could not have afforded to take this time off if it weren't for this program. So, thanks NIH.
OK - so some nuts and bolts. You get a health related doctoral degree (specifics on eligibility here). Graduate. Get a job doing research at a non-profit, preferably NIH funded (post-doc sort of thing). They pay off your loans! Each quarter I get notified that they sent $8,750 off to Sallie Mae. The only continuing work, besides the research, is that your supervisor has to fill out a form saying you are working hard (thanks +Alexander Anderson) and you have to verify that the money is going where it is supposed to. Easy.
Here are some links that the folks at NIHLRP provided me to supply with data about who has gotten funded.
http://www.lrp.nih.gov/pdf/LRPFY2012DataBook.pdf
http://www.lrp.nih.gov/pdf/1211.1_trends_report.pdf
The application process was similar to any other grant writing endeavor. There was a 6 page project description as well as a biosketch and some institution-specific paperwork. Importantly, a major component is the training program that you and your mentor create, which must be described in detail in a letter. For me, this included a PhD training program, but this is not necessary. Here is my successful grant application:
You can then, competitively, renew this grant as well. I don't know how that process goes since I missed the renewal application cycle (whoops!). It turns out, they don't remind you... But, I'll be submitting a renewal application this year, as long as there isn't a 5 year gap, you can renew. This is important, as when I go back to finish my clinical training, I will not qualify, as I won't be able to dedicate at least 50% of my time to the research. However, once I am faculty again, as long as I have educational debt and am spending 50% of my time doing qualifying research, I can continue to apply.
Long winded, sorry. Short story: great program. Give it a shot! We need more health professionals doing research, and this is a great way to defray some of the opportunity cost of not practicing.
At press time - I realize I've forgotten a few key points. They are:
Application cycle runs September 1-Nov 15th. Funding level is nearly 50%, so your odds are good, and the initial contract period is 2 years. Go apply!
Also - here is a 'tip sheet' on best practices for applications: http://www.lrp.nih.gov/pdf/
I distinctly remember one day during MS-1 when I was shadowing a hand surgeon in the OR at the Louis Stoke VA (our teaching VA in Cleveland), and the surgery resident, a PGY-2 in general surgery, described the NIH Loan Repayment plan to me. He was going to take a few years off after his PGY-3 year to pursue a PhD in tissue engineering. The benefit of doing this, he said, was that he could pick his clinical interest first, then pursue a related research area, rather than the MSTP folks who have to pick a scientific discipline before they know what sort of doctor they'll be. This has come into sharper focus for me as my best friend from medical school did his PhD in cardiac stem cells, assuming he would do cardiology, and fell in love with urologic surgery. It isn't so much that his training was wasted, it certainly wasn't, but he lost the momentum that he could have had had his clinical interests been aligned with his doctoral studies. So, in this sense, I'm glad that I wasn't accepted into the MSTP programs, as by going straight through medical school, I found my 'calling' as an oncologist, and then subsequently found how research fits into that - and this is an important distinction.
So, to continue the story, I chose radiation oncology, and now, between PGY-4 and 5, I have been taking time off to do dedicated translational research, and because of the NIH extramural loan repayment program, I am paying off a year of medical school for each year of qualified research! (Considering the opportunity cost of not practicing radiation oncology, I am still a bit 'behind' each year I postpone taking a faculty job, but this dulls the pain significantly.)
While I was disappointed that I didn't get in to the MSTP programs, I truly feel that, at least for me, this path is one that will better suit me for a rewarding career. Obviously, each person is different - had I had a burning passion about a certain kind of research early on, then following that path when I was younger might have been easier. I certainly face a whole slew of personal difficulties as a 37 year old PhD student with 2 young kids that I wouldn't have had I pursued this earlier. On the other side of the coin, with nearly ten years of medical experience, I bring a lot more to the table for my research than I would have after MS-2. At the end of the day, I am happy that the NIH funds both pathways, and we are in dire need of more physicians doing science. Like I said in my TEDMED talk, I truly feel that the physician-scientist plays a unique role in science. Indeed we are specifically trained to connect seemingly disparate pieces of data to build a theory, like a constellation of symptoms to build a diagnosis. As science gets more and more specialised, and knowledge silos get deeper and deeper, I feel this role will grow in importance as well - highlighting the need to keep these programs going.
While I love science and research, I came out of medical school with ~$200,000 in debt. I could not have afforded to take this time off if it weren't for this program. So, thanks NIH.
OK - so some nuts and bolts. You get a health related doctoral degree (specifics on eligibility here). Graduate. Get a job doing research at a non-profit, preferably NIH funded (post-doc sort of thing). They pay off your loans! Each quarter I get notified that they sent $8,750 off to Sallie Mae. The only continuing work, besides the research, is that your supervisor has to fill out a form saying you are working hard (thanks +Alexander Anderson) and you have to verify that the money is going where it is supposed to. Easy.
Here are some links that the folks at NIHLRP provided me to supply with data about who has gotten funded.
http://www.lrp.nih.gov/pdf/LRPFY2012DataBook.pdf
http://www.lrp.nih.gov/pdf/1211.1_trends_report.pdf
The application process was similar to any other grant writing endeavor. There was a 6 page project description as well as a biosketch and some institution-specific paperwork. Importantly, a major component is the training program that you and your mentor create, which must be described in detail in a letter. For me, this included a PhD training program, but this is not necessary. Here is my successful grant application:
You can then, competitively, renew this grant as well. I don't know how that process goes since I missed the renewal application cycle (whoops!). It turns out, they don't remind you... But, I'll be submitting a renewal application this year, as long as there isn't a 5 year gap, you can renew. This is important, as when I go back to finish my clinical training, I will not qualify, as I won't be able to dedicate at least 50% of my time to the research. However, once I am faculty again, as long as I have educational debt and am spending 50% of my time doing qualifying research, I can continue to apply.
Long winded, sorry. Short story: great program. Give it a shot! We need more health professionals doing research, and this is a great way to defray some of the opportunity cost of not practicing.
At press time - I realize I've forgotten a few key points. They are:
Application cycle runs September 1-Nov 15th. Funding level is nearly 50%, so your odds are good, and the initial contract period is 2 years. Go apply!
Also - here is a 'tip sheet' on best practices for applications: http://www.lrp.nih.gov/pdf/
Monday, June 3, 2013
Guest post day - The Tree of Life
A post from me is featured on the Tree of Life today trying to answer the question:
How do we build trust in the biological sciences to promote the use of preprints?
So head over and check it out.
http://phylogenomics.blogspot.com/2013/06/guest-post-from-jake-scott-building.html
There's also a new post up at Warburg's Lens discussing some work by +Benjamin Werner and Arne Traulsen at Max Planck.
Next up I'll post about the NIH Extramural Loan Repayment program.
How do we build trust in the biological sciences to promote the use of preprints?
So head over and check it out.
http://phylogenomics.blogspot.com/2013/06/guest-post-from-jake-scott-building.html
There's also a new post up at Warburg's Lens discussing some work by +Benjamin Werner and Arne Traulsen at Max Planck.
Next up I'll post about the NIH Extramural Loan Repayment program.
Subscribe to:
Posts (Atom)