As a clinician cum scientist, I am almost always involved in collaborations with multi-disciplinary groups. And, as an american who has done some training in the UK, many of those groups are NOWHERE REMOTELY NEAR one another. To keep myself, and my collaborators sane, I've been constantly experimenting with different combinations of cloud based repositories so that we can all work on things together.
So, I started out with +Dropbox by itself, but quickly found that, without spending money, I quickly filled up my allotted space. Even after totally maxing out the freebie upgrades (I think I have like 35Gb or something thanks to some promo at Oxford, referring everyone I know and being a shameless social media promoter - every Mb counts!), I still am limited. So I added +SugarSync for my personal files, to clear off dropbox and leave it for just collaboration. Well, that filled up pretty quick too, and as I haven't found sugarsync to be as easy to navigate as dropbox, I haven't worked as hard to maximize my space there. I muddled along with these two but still struggled with things like version control and the desire for contemporaneous editing. Which led to...
Google drive and google docs. So, with google drive, you get 25Gb or something, but I started to get confused as to where things were. When I wanted to collaborate real time though, I was stuck, as google docs was really the best thing going (and still is, for many things). However, in the last few months, I discovered +writeLaTeX, which has really changed the way I do business. This is a (mostly) free service (I haven't hit the wall yet, and it is pretty big considering that you don't store data there really) in which you can create, manage and edit LaTeX documents with as many people as you like. For me, as a mathematical biologist, this was great - so long as my collaborators were also theorists who were tex-savvy. I first used it doing a little hack-a-thon with some friends that I blogged about before. Recently though, as in, in the last few days, they have added a Rich Text formatting layer which, I think, will change everything again. No longer will I have to give the link to a document to my biological/clinical collaborator with the caveat 'just ignore everything that isn't text - squint a bit if you have to'. Now, they can just go ahead and edit away just like they are in word or whatever, but I can come in behind and have the full functionality of LaTeX. So that aspect seems solved.
**note - I LOVE writelatex, and you should check it out... and if you do, use this code and I get 50MB more space. https://www.writelatex.com/signup?ref=5ed6049a8e70 **
The only issue left for me is, now that I'm cranking TONS of simulations in (what I hope is) the final push toward this whole PhD thing, how to work on a DATA-HOG of a project on two computers (I can't stay at work late because I want to go home to my super sweet little kids, but then I DO want to work after they go to bed without driving back to work) or to share this with someone else. I thought, at first, that I'd just clear out my dropbox a bit, and try to be parsimonious with what I saved... but, as the output for my simulations is \mathcal{O} 30Gb/simulation, this quickly became unfeasible. Enter bittorrentsync - my saviour. This little gem (free) lets you have a synced drive on as many computers as you like. The freeing move here is that there is no cloud interaction, so there is NO SPACE LIMITATION. (There is also no auto-backup for the same reason... but this is obviated if you use Time Machine or something similar). All you do is download the software (a couple Mb's) and then, to set up a shared directory, it generates a 'secret' which you share with whomever you want to share. The secret is a massive string of letters/numbers that is autogenerated - and I bet even the guy at xkcd.com would approve.
So, now I'm set and I don't have to think too hard. I also don't forsee ever having to spend any money (until maybe I have a lab or my own and lots of people, but by then - if that time ever comes - I will have grant money to spend on such things... ?).
Summary: I use dropbox for shared/travelling talks, figures, syncing my papers library, shared simple code (matlab, etc). I use sugarsync for my personal documents (though I might phase this out...). I use writelatex for (now and going forward) ALL papers I write and bittorrent sync for shared working directories (also home/office syncing of working directories).
As an aside, dropbox has a cool feature - since everyone can modify things at will, they have a nice cloud backup system, which includes who the last person to interact with a document was, so in case your supervisor starts randomly deleting hunks of your thesis, you can call them on it - not naming any names... +Alexander Anderson :)
To be fair, I don't think Sandy actually deleted this stuff... and I was able to recover it, but it was pretty funny :)
Sunday, December 15, 2013
Monday, December 9, 2013
Guest post on the Oxford Centre for Maths Biology Blog
I wrote a post for my group's new blog on the importance of #preprints in science, and specifically on the new #bioRxiv. Head on over and check it out:
http://wcmb-oxford.blogspot.com/2013/12/pre-print-servers-dangerous-ways-to.html
http://wcmb-oxford.blogspot.com/2013/12/pre-print-servers-dangerous-ways-to.html
Thursday, December 5, 2013
Investigating the effects of microenvironmental perturbation on a stem driven tumor
I've been interested in the cancer stem cell hypothesis for some time, a subject that my colleague +Heiko Enderling has been thinking about and modeling for some time (list of his pubs here). I first became interested in this concept before I became a DPhil student and member of the Integrated Mathematical Oncology group, when I was a clinical resident in radiation oncology at the Moffitt Cancer Center. One of the first papers that I saw that truly scared me was a paper by Tamura and colleagues (abstract) that showed that recurrent glioblastomas had a significant increase in CD-133 staining (a stain commonly associated with stemness in this cancer and others), and that this increase is correlated with (maybe causes, jury yet out) an increase in aggressiveness of the recurrent tumor and a decrease in its sensitivity to treatment.

The standard rationale for the latter is that these special 'stem cells' have a higher intrinsic resistance to radiation therapy (which I don't argue), but I subsequently wrote down a series of simple (some might say Noddy) ODE models which suggested, at least to me, that there might also be a stem promoting effect of radiation. Since then, I have found that this effect has been shown in breast cancer, and further, that there have been a number of microenvironmental perturbations that have been shown to do the same thing, though only in a qualitative way - many of these articles have had my collaborator, +Anita Hjelmeland as a co-author, and I talked about them quite a bit in a previous blog post on our recent R-01 submission.
When I first started on this problem from the theoretical standpoint, it was with ODE models, as I mentioned. But since then, +David Basanta and +Alexander Anderson and I have worked to build a cellular automaton model of a stem-driven tumor which included blood vessels as sources of oxygen. We built this simple model to test if there were some sort of intrinsic/emergent change in the resultant tissue phenotype when the overall levels of oxygen supplied to it went down (when we reduced the density of the vessels). Below you can see the schematic of our model system. On the left is the canonical Cancer Stem Cell hypothesis (which I have major issues with... more to come in a few weeks I hope) and on the right there is our CA model rules.

After much simulation and effort we found... in short, that no, there is not. Our initial, negative results, were frustrating, but after discussions with +Anita Hjelmeland and Prakash Chinnaiyan (a biologist and clinician, respectively) we found that the model was telling us more... while we found little qualitative effect when we changed the vascular density, changing the instrinsic stem behavior parameters in the presence of a minimalistic environment revealed that there were only three major meta-phentypic behaviors possible: extinction, dormancy (homeostasis) and overgrowth as seen below.
Our frustration at not seeing the emergence of greater stem-fraction upon lowering the oxygen levels however, led to the rather obvious (in retrospect) conclusion that only through a modification of the symmetric division rate of the stem cells (modified by hypoxia) could this effect be recapitulated. Interestingly, this is a conclusion we had come to in the past using a simpler model system (ODEs) but it was never convincing (though there is a nice piece in J Theoretical Biology which gave me some confidence... here). Initial testing of this hypothesis, by modification of the CA rules to include this change are striking - surrounding the area of hypoxia, we have an emergent stem cell niche.
| Stem cells (red) emerge in areas surrounding necrosis (white/center) when the CA rules allow biased symmetric division in the presence of moderate hypoxia (right). |
We have just begun exploring this phenomenon, and indeed there is quite a bit of work to do, both theoretically and experimentally, to validate and better characterize what is going on - so that's why we wrote an R-01.
Anyways, if you want more details, you can read the paper on the bioRxiv now, or in a few days on PLoS Computational Biology where it has been accepted and is nearing publication. We've also made the baseline code freely available on sourceforge here.
If you have comments on the paper itself, please leave them on the bioRxiv site so anyone can see them (or wait to put them on the PLoS CB site). Ok, back to work.
Monday, October 21, 2013
A new pre-print server for biology: the bioRxiv
I've written a few posts in the past about the need for a pre-print server in biology, like the one that physics and maths have, in the arXiv, to speed up the rate of dissemination of information in science, and to help promote open access. As a physicist by original training, posting my work before it is accepted or done has never seemed like a big deal - it is science after all, we're all wrong, all the time (but we're trying to get closer to the truth, AS A GROUP).
In my community, the theoretical (mathematical) oncology community, we have been able to get around the lack of a standard pre-print server as there is a quant-bio section to the current arXiv, but it is certainly not widely read, nor is it really what the folks at the arXiv want to support (but we thank them for doing so, mind you). In fact, I once tried to get them to add a theoretical oncology section without success - prompting me to create Warburg's Lens - a discussion forum for pre-prints in math oncology.
With the advent of +PeerJ there has been at least one option, and another called CancerCommons has sprung up as well. In the next several weeks, we'll have another option, the bioRxiv - run by the folks at Cold Spring Harbor - a highly respected biological institute in New York. There have been a few attempts at this sort of thing before - Nature tried it once with its "preceedings", but it never took hold. I heard a rumor that this is because a lot of non-science was posted (thinly masked creationism and silly studies about herbal supplement pyramid schemes like Protandim). So, the bioRxiv has a plan to prevent this: they've asked a number of people to become "affiliates" whose job it is to screen the preprints to make sure that it is, at least, science. There will be no judgement about merit, we're to leave that to the communities, but just to screen out non-science. Anywho, I'll be splitting my pre-print posts between here and the physics arXiv from now on - depending on focus. More clinical/biological papers will go to the bioRxiv, and more mathematical/methodological will go to the physics arXiv. For Warburg's Lens, I'll troll both...
I hope you check out the bioRxiv, and, if you are doing work in the biological sciences, I hope you consider posting your work here as you submit to standard journals. I did a poll recently on a friend's website, and found that the only thing really stopping biologists from posting was... well... NOTHING. Mostly, it was habit and inertia. So - let's change those. Let's put our work out there early and often and let the scientific community do its thing!
In my community, the theoretical (mathematical) oncology community, we have been able to get around the lack of a standard pre-print server as there is a quant-bio section to the current arXiv, but it is certainly not widely read, nor is it really what the folks at the arXiv want to support (but we thank them for doing so, mind you). In fact, I once tried to get them to add a theoretical oncology section without success - prompting me to create Warburg's Lens - a discussion forum for pre-prints in math oncology.
With the advent of +PeerJ there has been at least one option, and another called CancerCommons has sprung up as well. In the next several weeks, we'll have another option, the bioRxiv - run by the folks at Cold Spring Harbor - a highly respected biological institute in New York. There have been a few attempts at this sort of thing before - Nature tried it once with its "preceedings", but it never took hold. I heard a rumor that this is because a lot of non-science was posted (thinly masked creationism and silly studies about herbal supplement pyramid schemes like Protandim). So, the bioRxiv has a plan to prevent this: they've asked a number of people to become "affiliates" whose job it is to screen the preprints to make sure that it is, at least, science. There will be no judgement about merit, we're to leave that to the communities, but just to screen out non-science. Anywho, I'll be splitting my pre-print posts between here and the physics arXiv from now on - depending on focus. More clinical/biological papers will go to the bioRxiv, and more mathematical/methodological will go to the physics arXiv. For Warburg's Lens, I'll troll both...
I hope you check out the bioRxiv, and, if you are doing work in the biological sciences, I hope you consider posting your work here as you submit to standard journals. I did a poll recently on a friend's website, and found that the only thing really stopping biologists from posting was... well... NOTHING. Mostly, it was habit and inertia. So - let's change those. Let's put our work out there early and often and let the scientific community do its thing!
Tuesday, October 8, 2013
Glioblastoma: Stem cells, plasticity and the niche(s). A summary and introduction to our R-01 submission.
I've had an interest, both clinically and scientifically, in glioblastoma for about 6 years. This started out because I was given a project as a medical student looking at outcomes of treatment for elderly patients, but has continued and consumed more and more of my conscious and unconscious thought in the intervening years. For a long time, patients over 70 weren't given the same care as their younger counterparts because it was felt that the treatment was too harsh for them. This paradigm has begun to change, thanks in some part to the work I was a part of, but the outcomes remain very poor, for all patients. While I had success in this initial clinical research (first two figs), I was frustrated at what I perceived as a lack of progress, and didn't feel that I was really contributing to this. This feeling is really what pushed me into basic research...
So, our standard of care is to treat where the tumor was before surgery and a smallish margin around the edges, and hoping that our chemotherapy will take care of the more distant cells. In 2004, a famous paper from Singh and colleagues identified a small subset of cells within a glioblastoma which seemed to be responsible for these recurrences - and these cells shared many attributes of non-cancer stem cells. It turned out that these cells were more resistant to radiation - and this gave us hope: maybe we weren't curing these patients because we were targeting the wrong cells! The field of glioma stem cell biology has advanced rapidly and many scientists are working on the problem. A prominent group in this field is led by Jeremy Rich at the Cleveland Clinic's Lerner Research Institute. They have made many advances, but recently their focus has been on the effect of physical/chemical factors within the 'microenvironment' of the tumor that promote these special 'stem' cells - for example acidity, low oxygen tension and low glucose levels.
One of the scientists from this lab in Cleveland, Anita Hjelmeland, recently took a faculty position at the University of Alabama in Birmingham, where they have a massive brain tumor center (called a SPORE) to continue her work. She and I met when I was visiting her lab in Cleveland and gave a talk about some of the theory work that +David Basanta and I have done. She and I hit it off, scientifically, and we decided to start a collaboration. We've worked together now on a few different projects that are in various phases of development, and just yesterday, we submitted an R-01 (a large scale, 5 year grant) proposal that is led by Anita and David, with participation from myself, +Alexander Anderson and +Heiko Enderling. We are hoping to leverage the strengths of her biological laboratory with our theoretical modeling techniques to try to make some progress against this cancer. Her work previously has shown, convincingly, that hyoxia (low oxygen levels), acidic pH and low glucose levels promote these 'stem' cells, but trying to understand how they all work together is a difficult task - especially in a living system. This is where our computational models can help.
Anyways, we have our fingers crossed for the success of our grant, and I'll be sure to keep you updated as to our progress. As a teaser, I am including a figure from our grant. Mind you, this is unpublished preliminary data (SHARING IN SCIENCE IS GOOD!), so don't draw too many conclusions from it.
Some background: it seems that these special 'stem' cells in the tumor preferentially live in special areas called 'niches'. These niches come in (at least) two different varieties, near to the vasculature, where nutrients are abundant, and near to areas of necrosis (cell death) where nutrients are scarce. This difference is intriguing and some observations have suggested that they might contribute to differences in treatment response. So - the preliminary finding... our model suggests that the physical microenvironmental history of these niches is very different, and that the evolutionary dynamics within them are as well! If we can better understand how these niches are created and maintained, maybe we can make some inroads against the progression of this tumor.
So, our standard of care is to treat where the tumor was before surgery and a smallish margin around the edges, and hoping that our chemotherapy will take care of the more distant cells. In 2004, a famous paper from Singh and colleagues identified a small subset of cells within a glioblastoma which seemed to be responsible for these recurrences - and these cells shared many attributes of non-cancer stem cells. It turned out that these cells were more resistant to radiation - and this gave us hope: maybe we weren't curing these patients because we were targeting the wrong cells! The field of glioma stem cell biology has advanced rapidly and many scientists are working on the problem. A prominent group in this field is led by Jeremy Rich at the Cleveland Clinic's Lerner Research Institute. They have made many advances, but recently their focus has been on the effect of physical/chemical factors within the 'microenvironment' of the tumor that promote these special 'stem' cells - for example acidity, low oxygen tension and low glucose levels.
One of the scientists from this lab in Cleveland, Anita Hjelmeland, recently took a faculty position at the University of Alabama in Birmingham, where they have a massive brain tumor center (called a SPORE) to continue her work. She and I met when I was visiting her lab in Cleveland and gave a talk about some of the theory work that +David Basanta and I have done. She and I hit it off, scientifically, and we decided to start a collaboration. We've worked together now on a few different projects that are in various phases of development, and just yesterday, we submitted an R-01 (a large scale, 5 year grant) proposal that is led by Anita and David, with participation from myself, +Alexander Anderson and +Heiko Enderling. We are hoping to leverage the strengths of her biological laboratory with our theoretical modeling techniques to try to make some progress against this cancer. Her work previously has shown, convincingly, that hyoxia (low oxygen levels), acidic pH and low glucose levels promote these 'stem' cells, but trying to understand how they all work together is a difficult task - especially in a living system. This is where our computational models can help.
Anyways, we have our fingers crossed for the success of our grant, and I'll be sure to keep you updated as to our progress. As a teaser, I am including a figure from our grant. Mind you, this is unpublished preliminary data (SHARING IN SCIENCE IS GOOD!), so don't draw too many conclusions from it.
Some background: it seems that these special 'stem' cells in the tumor preferentially live in special areas called 'niches'. These niches come in (at least) two different varieties, near to the vasculature, where nutrients are abundant, and near to areas of necrosis (cell death) where nutrients are scarce. This difference is intriguing and some observations have suggested that they might contribute to differences in treatment response. So - the preliminary finding... our model suggests that the physical microenvironmental history of these niches is very different, and that the evolutionary dynamics within them are as well! If we can better understand how these niches are created and maintained, maybe we can make some inroads against the progression of this tumor.
So - wish us luck. The model that we have extended to make the above prediction is under review at PLoS Computational Biology, but you can see a preprint here. Anita has published a TON of papers on this subject, which you can find on pubmed. David has also written several papers (a couple with me) on glioblastoma, but only the preprint so far on stem cells in this disease. Our resident stem cell modeling expert, +Heiko Enderling, will be of great help as well. You can see his long publication record on his website.
Tuesday, September 17, 2013
Pint of Science, US
Wow - it's been a while since I wrote anything. A combination of several academic visits from +Alex Fletcher and Anita Hjelmeland, trying desperately to make some headway on my thesis, and a teething toddler left this blogging effort at the bottom of the pile. I've got a few posts I need to write to catch up, but I thought I'd start with this one.
Last year, a movement was started in the UK called pint of science who's stated mission is to bring top scientist into pubs to communicate their passion for science to everyman. They began with 15 pubs in London, Oxford and Cambridge for a several days long festival to great acclaim. The movement is now spreading, and my good friends +Parmvir Bahia, +David Basanta, +Arturo Araujo and new friend +Angela Rey decided to start the movement here in the US - aptly named, Pint of Science, US. They have worked to find their own spin on the theme, and settled on holding both a live event, and also on a podcast, on a monthly basis. The central theme is the same - communicate what it is that we do as scientists, and why we do it, to the folks with whom we live and work.
The first even was held two weeks ago today at the New World Brewery in Tampa, FL. I was honored to be asked by my friends to be the first speaker. This is both a good thing and a bad thing. The expectations are low (or, undefined), but it is also a difficult ask, because there is no predetermined script to follow. So, you'll notice some hems and hahs in my podcast.
The audience at this, the inaugural event, consisted entirely of friends and co-workers, so it felt a little strange telling my life story and trying to convince 'people' that mathematics can help to play a role in cancer research (as everyone knew my life story and DOES mathematical cancer research). So, in some ways, this event was more difficult than any other event like this that I've done. Strange to admit, but I was sweating bullets!
Last year, a movement was started in the UK called pint of science who's stated mission is to bring top scientist into pubs to communicate their passion for science to everyman. They began with 15 pubs in London, Oxford and Cambridge for a several days long festival to great acclaim. The movement is now spreading, and my good friends +Parmvir Bahia, +David Basanta, +Arturo Araujo and new friend +Angela Rey decided to start the movement here in the US - aptly named, Pint of Science, US. They have worked to find their own spin on the theme, and settled on holding both a live event, and also on a podcast, on a monthly basis. The central theme is the same - communicate what it is that we do as scientists, and why we do it, to the folks with whom we live and work.
The first even was held two weeks ago today at the New World Brewery in Tampa, FL. I was honored to be asked by my friends to be the first speaker. This is both a good thing and a bad thing. The expectations are low (or, undefined), but it is also a difficult ask, because there is no predetermined script to follow. So, you'll notice some hems and hahs in my podcast.
 |
| PB doing the introduction... |
The audience at this, the inaugural event, consisted entirely of friends and co-workers, so it felt a little strange telling my life story and trying to convince 'people' that mathematics can help to play a role in cancer research (as everyone knew my life story and DOES mathematical cancer research). So, in some ways, this event was more difficult than any other event like this that I've done. Strange to admit, but I was sweating bullets!
Anywho, after some gentle editing from the Pint of Science US team, a podcast emerged and the event was a success. I had a great time participating, and look forward to the next even when I can have a pint and learn about some neuroscience from our friend Tom Taylor-Clark, at USF.
I certainly recommend coming to the event, and subscribing to the podcasts - which are available on iTunes as well as through the link here. You can also follow Pint of Science US on G+ or on twitter @pintofscienceUS for updates.
Have a pint, talk about science.
Monday, July 29, 2013
A visitor, the resulting hackathon, and a nice result.
So, I was sitting in a pub in Oxford (the head of the River - gorgeous place and one that Lewis frequents), and I met this guy +Artem Kaznatcheev. No, he wasn't having a pint at the table next to me. No, he wasn't there for an academic visit. I met him on twitter, because of a tweet from +Steven Strogatz about mathematics in biology. Here's how it all began:
In there, we can also see the first thoughts about making this blog - about 6 months before I actually did it. Better late than never?
Anyways, +Artem Kaznatcheev and +David Basanta and I (and some others) started what is now a 10 month long conversation, mostly on Google+, in a community Artem started and we co-moderate, called Evolutionary Game Theory. This conversation has covered topics (subsequently blogged about) ranging from understanding vs. predicting (followed up nicely by +Philip Gerlee in his blog here), games bacteria play (and a recent +Jeff Gore paper in PLoS Biology), connectors in science and, more recently, the topic of our original connection: the use of game theory in cancer.
The conversations have been lots of fun, we all think a bit differently, but have many of the same ideals about science, understanding and the uses of mathematics. Further, we are all hopeless nerds and *cough* workaholics. So, when Artem noticed that the conference he was going to (Swarmfest 2013) was near us, he jumped at the chance to come meet us and get some work done. On his way down, he gave a couple of David's papers a detailed read through to get the lay of the land (and blogged about it - clever way to annotate things for yourself as well as manage a post).
So, here's the hackathon part. Artem arrived Wednesday night and he and David worked into the wee hours. He then came in to #IMO and they spent the day doing a full analytic treatment of a game David and I published in the British Journal of Cancer. They then worked again into the wee hours at David's house. I arrived from an out of town trip the next day (Friday). Artem came in to IMO and gave a talk (which we managed to broadcast on G+, something we hope to continue, but with better sound quality, any ideas on a bluetooth mic?).
After his talk, we spent the afternoon identifying a tight question: in a growing tomour, what would change in a simple game or proliferative vs. motile cells between the middle and the edge, if anything?
One of the difficulties in EGT is that neighborhoods and population structure is not considered, indeed, it is assumed that the population is inviscid (well mixed). A great paper from Martin Nowak at Harvard gave us a way to think about effective neighborhood sizes (formally, how to understand changing game dynamics on graphs of differing, but regular, degree). This has some obvious applications to growing tumours - when they hit a basement membrane or an organ capsule they go from growing in 'free 3-d space' (neighbors on all sides) to growing almost in 2-d, against a wall (with neighbors only on one 'side').
So, we spent the rest of friday afternoon doing some analysis


After this experience, we hope to weave hackathons like this into our schedule more often. It certainly isn't something I (or my family) could tolerate every week, but it was fun and highly productive and we'll try to do it again soon.
Anyways, we'd love feedback on the paper. Artem has a more technical post today about the work and some future directions which you can read here.
In there, we can also see the first thoughts about making this blog - about 6 months before I actually did it. Better late than never?
Anyways, +Artem Kaznatcheev and +David Basanta and I (and some others) started what is now a 10 month long conversation, mostly on Google+, in a community Artem started and we co-moderate, called Evolutionary Game Theory. This conversation has covered topics (subsequently blogged about) ranging from understanding vs. predicting (followed up nicely by +Philip Gerlee in his blog here), games bacteria play (and a recent +Jeff Gore paper in PLoS Biology), connectors in science and, more recently, the topic of our original connection: the use of game theory in cancer.
The conversations have been lots of fun, we all think a bit differently, but have many of the same ideals about science, understanding and the uses of mathematics. Further, we are all hopeless nerds and *cough* workaholics. So, when Artem noticed that the conference he was going to (Swarmfest 2013) was near us, he jumped at the chance to come meet us and get some work done. On his way down, he gave a couple of David's papers a detailed read through to get the lay of the land (and blogged about it - clever way to annotate things for yourself as well as manage a post).
So, here's the hackathon part. Artem arrived Wednesday night and he and David worked into the wee hours. He then came in to #IMO and they spent the day doing a full analytic treatment of a game David and I published in the British Journal of Cancer. They then worked again into the wee hours at David's house. I arrived from an out of town trip the next day (Friday). Artem came in to IMO and gave a talk (which we managed to broadcast on G+, something we hope to continue, but with better sound quality, any ideas on a bluetooth mic?).
 |
| Here's me and Chandler looking interested. Also, you can see we had 4 or 5 others from all over, Germany, Oxford and I don't know where else... |
One of the difficulties in EGT is that neighborhoods and population structure is not considered, indeed, it is assumed that the population is inviscid (well mixed). A great paper from Martin Nowak at Harvard gave us a way to think about effective neighborhood sizes (formally, how to understand changing game dynamics on graphs of differing, but regular, degree). This has some obvious applications to growing tumours - when they hit a basement membrane or an organ capsule they go from growing in 'free 3-d space' (neighbors on all sides) to growing almost in 2-d, against a wall (with neighbors only on one 'side').
So, we spent the rest of friday afternoon doing some analysis


Then, on Friday night, we celebrated by buying a bunch of redbulls and working 'till 2am at my house. I dropped him off at his hotel, then picked him up for a late breakfast and we worked, using this great new on-line app we found +writeLaTeX (which is AWESOME) and started a manuscript. At dinner time, we broke company... Then, Saturday night, we really blew off some steam - and made the figures for the paper. Dropped off at his hotel around 2am, he was picked up by David the next morning and they worked until his plane left.
So, that was the hackathon. The result, we are proud to announce, is a paper, done and dusted, beginning to end, in 15 days, with the lion's share of the work done in the first 4 days (about 48 hours of which saw the three of us working full on). To be fair, we thought hard about the question in conversations for several months, and the groundwork had been laid by previous papers, but really, this felt like doing theory the way you're meant to. It felt inspired. And, I think, this is the best paper I've been a part of so far. But, don't take my word for it, check out the preprint, just released on the arXiv today:
We also have submitted it, and it is now under consideration at the Proceedings of the Royal Society, Series B. While we were motivated by a cancer scenario, we feel the result applies more broadly to biology than many of our previous papers, and so have targeted a broader biological journal. And, PRS B does publish theory, and has recently published some interesting work from +Arne Traulsen's group at Max Planck on evolution in structured populations, so it seemed like we have a chance... we'll see.
Anyways, we'd love feedback on the paper. Artem has a more technical post today about the work and some future directions which you can read here.
Friday, July 19, 2013
Visit to Summer Science Program in Socorro, NM
About a year and a half ago, after my talk at TEDMED 2012, I got a call from a medical oncologist asking if I would come to visit a summer camp in the desert of New Mexico that featured a bunch of really smart kids and astrophysics. This sounded right up my alley (I love nerds, and I love stars), but the trip from Oxford to New Mexico seemed a little bit... far. So, I said I was interested, but maybe we could talk next year, when I was back in Tampa in IMO - which I promptly forgot about. Thankfully, they didn't forget, and a year later, I got the call again. So I packed up and headed for the Summer Science Program's, Socorro, New Mexico campus (on the campus of New Mexico tech, home of the Miners and well known for excellent work on explosives!).
As I left the rental car place in the Albuquerque airport, I asked the attendant how to get to 25 South and he said:
"You mean 25 North, there isn't anything to the south"
Which is how I knew it was the right direction...
After arriving, I was met at my hotel by the site director, Barb, and I asked to go up to the telescope to see the kids doing their observations. This camp is set up in a really cool way, they spend 6 hours a day in the classroom doing intensive math/physics and computer science (Python) coursework. They are split into teams of three, and at the very beginning they choose a near earth asteroid (the way they choose the asteroids is kind of a funny story, but maybe one of the students will comment with how that works...). The teams then get a certain amount of telescope time, during which they take measurements of their asteroid's position. They are then expected to do the math and write up some code to predict the orbit - which they then share with some astrophysicists at Harvard. Really cool - not just taking courses and playing with telescopes, but DOING MEANINGFUL SCIENCE. Needless to say, I didn't fly all the way to NM to just give a talk, so at around midnight, I wandered up to the observatory. It was DARK (perfect) and Barb got out a flashlight, which I asked her to extinguish so I could enjoy the darkness... she demurred suggesting that we needed it to see any snakes in the path.
Me: Snakes, pshaw... wait, you mean like that one?
She ran away, and I snapped this pic - can anyone ID it? Not the best pic - iPhone flash sucks... its head was narrow, like a non-venomous snake, but I don't know my high desert fauna... little help?
We finally got to the telescope and found the group observing.
They weren't able to see their asteroid this night, but they showed me a beautiful globular cluster (pictured) and a spiral galaxy that they found. How cool.
The TA who was there gave me a short tour and I found out that he is starting his DPhil in Oxford next year, at Summerville college (right next to the CMB, my home!) doing condensed matter physics. Small world.
I finally crashed and awoke to take a short run and saw this really cool 'M' in the hills. This delighted my daughter (Maren - of course the 'M' was for Maren!) who thought of the Thomas the tank engine episode about the Man in the hills. It turns out it is for the NM tech Miners, but I like the M in the hills better.
Anyways, after my run, I went up to the campus and set up to give my talk,.which you can see here:
Beyond an introduction into the uses of mathematics in cancer research (and theoretical biology in general) I focused on the need for taking risks in science, and how we ought not shy away from creative thinking. Further, I talked a bit about my tortuous career path and how having done a ton of different things, and not just following a straight arrow course, had informed my science and my life in general. After the talk, they gave me a sweet green SSP engraved laser pointer (THANKS!) and I had lunch with the students and a chat afterwards... it was at this point that I started ranting about open access science (I was overtired) and tried to convince them to put the findings from their summer research onto the arXiv (might need an endorsement for astrophysics...anyone willing to help?) and their asteroid tracking code onto github. Why not!?
Here's the pic. I was overjoyed to see a large cadre of girls at the camp - a good sign for our future! I am certainly going to keep this place, a well-kept secret, in mind for when my kiddos are in high school. I just wish I had had an opportunity like this - and I'm amazed that I had never heard of it. My high school teacher - Bob Shurtz (LEGEND) - is the coach of the US Physics Olympiad team and is well plugged in in these matters, but had never heard of this camp. Considering it has been around since Sputnik, this surprised me. Further, there were kids from all over the world - India, Hungary and China in addition to the US. Oh well. In my next lifetime...
Oh yeah... it was HOT.
Also worth checking out - the students at SSP have a blog. Good stuff. Watch these kids - they are a bright crew. And, with any luck, I interested one of two of them in theoretical biology!
Great experience as a lecturer, looks amazing as a student. Spread the word.
As I left the rental car place in the Albuquerque airport, I asked the attendant how to get to 25 South and he said:
"You mean 25 North, there isn't anything to the south"
Which is how I knew it was the right direction...
After arriving, I was met at my hotel by the site director, Barb, and I asked to go up to the telescope to see the kids doing their observations. This camp is set up in a really cool way, they spend 6 hours a day in the classroom doing intensive math/physics and computer science (Python) coursework. They are split into teams of three, and at the very beginning they choose a near earth asteroid (the way they choose the asteroids is kind of a funny story, but maybe one of the students will comment with how that works...). The teams then get a certain amount of telescope time, during which they take measurements of their asteroid's position. They are then expected to do the math and write up some code to predict the orbit - which they then share with some astrophysicists at Harvard. Really cool - not just taking courses and playing with telescopes, but DOING MEANINGFUL SCIENCE. Needless to say, I didn't fly all the way to NM to just give a talk, so at around midnight, I wandered up to the observatory. It was DARK (perfect) and Barb got out a flashlight, which I asked her to extinguish so I could enjoy the darkness... she demurred suggesting that we needed it to see any snakes in the path.
Me: Snakes, pshaw... wait, you mean like that one?
She ran away, and I snapped this pic - can anyone ID it? Not the best pic - iPhone flash sucks... its head was narrow, like a non-venomous snake, but I don't know my high desert fauna... little help?
We finally got to the telescope and found the group observing.
They weren't able to see their asteroid this night, but they showed me a beautiful globular cluster (pictured) and a spiral galaxy that they found. How cool.
The TA who was there gave me a short tour and I found out that he is starting his DPhil in Oxford next year, at Summerville college (right next to the CMB, my home!) doing condensed matter physics. Small world.
I finally crashed and awoke to take a short run and saw this really cool 'M' in the hills. This delighted my daughter (Maren - of course the 'M' was for Maren!) who thought of the Thomas the tank engine episode about the Man in the hills. It turns out it is for the NM tech Miners, but I like the M in the hills better.
Anyways, after my run, I went up to the campus and set up to give my talk,.which you can see here:
Beyond an introduction into the uses of mathematics in cancer research (and theoretical biology in general) I focused on the need for taking risks in science, and how we ought not shy away from creative thinking. Further, I talked a bit about my tortuous career path and how having done a ton of different things, and not just following a straight arrow course, had informed my science and my life in general. After the talk, they gave me a sweet green SSP engraved laser pointer (THANKS!) and I had lunch with the students and a chat afterwards... it was at this point that I started ranting about open access science (I was overtired) and tried to convince them to put the findings from their summer research onto the arXiv (might need an endorsement for astrophysics...anyone willing to help?) and their asteroid tracking code onto github. Why not!?
Here's the pic. I was overjoyed to see a large cadre of girls at the camp - a good sign for our future! I am certainly going to keep this place, a well-kept secret, in mind for when my kiddos are in high school. I just wish I had had an opportunity like this - and I'm amazed that I had never heard of it. My high school teacher - Bob Shurtz (LEGEND) - is the coach of the US Physics Olympiad team and is well plugged in in these matters, but had never heard of this camp. Considering it has been around since Sputnik, this surprised me. Further, there were kids from all over the world - India, Hungary and China in addition to the US. Oh well. In my next lifetime...
 |
| Yeah, that says 104F |
Also worth checking out - the students at SSP have a blog. Good stuff. Watch these kids - they are a bright crew. And, with any luck, I interested one of two of them in theoretical biology!
Great experience as a lecturer, looks amazing as a student. Spread the word.
Wednesday, July 3, 2013
The pre-metastatic niche is only half of the story of metastasis (it's the biological one)
Recently, Cancer Research UK posted an article on their blog in which they explain, in layman's terms, recent trends and ideas in research into metastatic spread. The focus of that article is on the concept of a 'pre-metastatic niche', the idea that the primary tumour emits signalling molecules that prime certain organs for the arrival of metastatic cells. We find this line of thought very interesting, as it could, at least in part, explain patterns of metastatic spread, but have strong opinions about how the ideas were presented and the lack of acknowledgment of the other factors that could be at play.
First, the reader is given a condensed historical background, in which the surgeon Stephen Paget is given credit for having solved the riddle of metastatic patterns 150 years ago. His method of studying metastatic spread in breast cancer is briefly mentioned, however, as is often the case when the seed-soil hypothesis is mentioned, these old 'truths' do not seem to be carefully checked. For example, a much more recent study from by Dr. J Pickren (reported in The Principles of Metastasis by L. Weiss, p. 231, recently reviewed here), which reports a 4:1 ratio between splenic and hepatic metastases (compared to the 14:1 ratio that Paget observed). Another fact not accounted for by Paget in his analysis, is that the liver not only receives arterial blood, but also blood from the gut organs via the portal vein, thereby increasing the chance of it receiving circulating tumour cells (CTCs). If micro-metasases are present in the gut, then these secondary CTCs will most likely lodge in the liver increasing the risk of developing liver metastases. Lastly, Paget only studied a single location of primary tumours, making general conclusions difficult to draw - especially as the connectivity differs greatly between organs. These simple observations should make it clear that Paget's hypothesis is nothing more than an indication of what might be the case in certain circumstances, rather than a settled fact.
From reading the article one also gets the impression that CTCs are drawn to certain organs in the body (e.g. the caption of the 2nd figure reading "Tumour cells are selective about where they end up." or later in text "...which wandering tumour cells find irresistible."). This is not in agreement with what we know today (and have known for the last 30 years) about the dynamics of metastasis formation.
On the contrary CTCs have little influence over where they end up, instead the correct picture is that of a primary tumour releasing astronomical numbers of CTCs into the blood stream (roughly 100 million cells per day, of which most die in the blood stream), and that these cells are distributed according to physiology of the circulatory system.
This means that each organ (except the lung and liver) receive a fraction of CTCs in direct relation to their relative blood supply, and only at this point, at which the cancer cells flow through the capillary bed of the organ, can organ specific mechanisms influence the fate of the cancer cell. This means that any explanation of why patterns of metastatic spread look as they do needs to first take into account the characteristics of the circulatory system, and only then the organ specific mechanisms such as the formation of a pre-metastatic niche.
These facts suggest (at least to us) that one should view the formation of the pre metastatic niche from a more passive point of view. The signals secreted by the primary tumour induce a systemic inflammatory response - which may or may not effect all organs. The evidence suggests that some distant sites respond in a way that makes them more hospitable to the CTCs that happen to pass though them and hence these cells are more likely to form overt metastases - but to present this as an active process is to stretch the data and to anthropomorphize to a dangerous extent.
When attempting to synthesize and communicate difficult scientific information to the public, it is always tempting to present a small slice of the story - and indeed, this is good practice as only so much can be communicated effectively at one time. But when doing this, it is essential to point out where the limits of our understanding are, and not oversell current hypotheses as the 'truth'. Science is, and always has been, a steady progression toward understanding, paved by models that are (we hope) less and less wrong. The way we think today is not likely to be the same as the way we think in 10 years time.
When attempting to synthesize and communicate difficult scientific information to the public, it is always tempting to present a small slice of the story - and indeed, this is good practice as only so much can be communicated effectively at one time. But when doing this, it is essential to point out where the limits of our understanding are, and not oversell current hypotheses as the 'truth'. Science is, and always has been, a steady progression toward understanding, paved by models that are (we hope) less and less wrong. The way we think today is not likely to be the same as the way we think in 10 years time.
Monday, June 17, 2013
Barriers to use of preprint servers
A week or two ago, I wrote a guest post on my friend +Jonathan Eisen's blog where I outlined what I thought were some problems standing in the way of folks utilizing #openaccess #preprint servers. I argued the micro-community building, like we're trying to do at Warburg's Lens, or that the folks at Haldane's Sieve have been doing, was a way forward. In that same post, I put a small survey, from which I learned quite a bit, and I want to share those results with you here.
I've never used google speadsheets before, so I learned some lessons about coding answers (doesn't deal well with age ranges etc.) and I had to do a BIT of recoding. There are lots of ways to look at the data, but I picked two to break out specifically. First some summary statistics though:
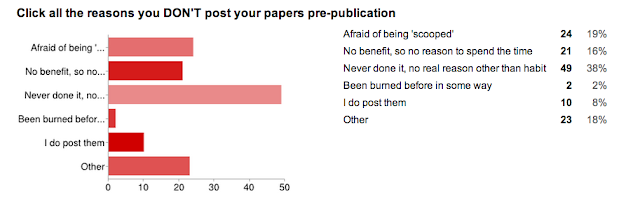
And... the reason we did the whole survey. WHY people don't utilize preprint servers.
I was really surprised by this one. I thought that the 'afraid of being scooped' answer would win the day (this is the case from my anecdotal experience with asking colleagues). I have a good answer to this one, to try to allay fears... but we don't even need it! The number one reason was:
"Pre-print servers? Meh."
And this is easy to change, it is just a culture shift, and one that is, I think in flux (in the right direction!).
You can see all the responses here, if you want, which includes all of the free text responses - where I think the real fodder for change lies. These are hard (impossible) to display, except to copy and paste them all in. But, I can tell you, having read them all, they fall into several categories:
- There is no benefit to me, so why spend the time
*** There is huge benefit, especially to early career folks, as you can show off your work to anyone, with just a link. Further, with the growing popularity of pre-print discussion forums, you can get valuable feedback before submission, hopefully making the process smoother.
- I want to, but my supervisor/collaborator won't let me.
*** Point those folks to these blog posts! Start the discussion.
- I would, if only some big fish in my field did it.
*** The Eisen brothers do it, so can you!
- I'd do it if tenure committees/funding agencies would give me credit.
*** This is a big issue, and I can only hope that the work we do here helps makes that change a reality.
- I'd do it if people could cite my preprints
*** THEY CAN!!! And, google scholar picks up those citations and counts them. Oh, brave new world, with such alt-metrics in it.
- I don't know the rules of the journals that I want to submit to, and I don't want to jeopardize my chances at "real" publication.
*** Just ask Wikipedia - here's the answer
- There's too many choices!
- There's too few choices!
Anyways, please do take a moment to read the responses, they are enlightening. I, for one, have learned that the barriers to getting pre-prints servers more utilized is simply one of outreach - so here I go. If you are in favor of speeding up (and opening up) science, take a minute to engage a scientist in your lab on this issue. It will pay dividends. There are plenty of options for preprint servers now, nicely outlined by +Jonathan Eisen in a post he wrote here, and even more coming on line. I am particularly excited about the bioRxiv coming out of Cold Springs Harbor, and am one of their affiliates in getting going. Until then, I have personally used the physics arXiv for my last five papers as well as a test use of PeerJ which has nice alt-metrics.
Ah, the paradox of choice.
Whatever you choose, post your preprints before you submit. Share your science. Whatever you study, let everyone else see stand on your shoulders so they can see farther, and further the cause.
I've never used google speadsheets before, so I learned some lessons about coding answers (doesn't deal well with age ranges etc.) and I had to do a BIT of recoding. There are lots of ways to look at the data, but I picked two to break out specifically. First some summary statistics though:
And... the reason we did the whole survey. WHY people don't utilize preprint servers.
I was really surprised by this one. I thought that the 'afraid of being scooped' answer would win the day (this is the case from my anecdotal experience with asking colleagues). I have a good answer to this one, to try to allay fears... but we don't even need it! The number one reason was:
"Pre-print servers? Meh."
And this is easy to change, it is just a culture shift, and one that is, I think in flux (in the right direction!).
You can see all the responses here, if you want, which includes all of the free text responses - where I think the real fodder for change lies. These are hard (impossible) to display, except to copy and paste them all in. But, I can tell you, having read them all, they fall into several categories:
- There is no benefit to me, so why spend the time
*** There is huge benefit, especially to early career folks, as you can show off your work to anyone, with just a link. Further, with the growing popularity of pre-print discussion forums, you can get valuable feedback before submission, hopefully making the process smoother.
- I want to, but my supervisor/collaborator won't let me.
*** Point those folks to these blog posts! Start the discussion.
- I would, if only some big fish in my field did it.
*** The Eisen brothers do it, so can you!
- I'd do it if tenure committees/funding agencies would give me credit.
*** This is a big issue, and I can only hope that the work we do here helps makes that change a reality.
- I'd do it if people could cite my preprints
*** THEY CAN!!! And, google scholar picks up those citations and counts them. Oh, brave new world, with such alt-metrics in it.
- I don't know the rules of the journals that I want to submit to, and I don't want to jeopardize my chances at "real" publication.
*** Just ask Wikipedia - here's the answer
- There's too many choices!
- There's too few choices!
Anyways, please do take a moment to read the responses, they are enlightening. I, for one, have learned that the barriers to getting pre-prints servers more utilized is simply one of outreach - so here I go. If you are in favor of speeding up (and opening up) science, take a minute to engage a scientist in your lab on this issue. It will pay dividends. There are plenty of options for preprint servers now, nicely outlined by +Jonathan Eisen in a post he wrote here, and even more coming on line. I am particularly excited about the bioRxiv coming out of Cold Springs Harbor, and am one of their affiliates in getting going. Until then, I have personally used the physics arXiv for my last five papers as well as a test use of PeerJ which has nice alt-metrics.
Ah, the paradox of choice.
Whatever you choose, post your preprints before you submit. Share your science. Whatever you study, let everyone else see stand on your shoulders so they can see farther, and further the cause.
Saturday, June 15, 2013
Institute for the Future: Health Horizons storification
I was an invited panelist at this year's Institute for the Future: Health Horizons meeting this past week. I had NO IDEA what I was getting myself into when I accepted the invitation, and I have to say, I was skeptical. My experience though, was without question, excellent. Quality organization, quality program, great attendees and a top-notch location. I'll write a proper post to describe the meeting in my own words, but until then, here is a storify version, in tweets. Most are my own, but I used some tweets from others when I was on stage, and also to fill in some spots where I slacked off!
Thanks a million to +Miriam Avery for the invite, and to all my other new friends at the IFTF. I hope to see you all again soon.
Friday, June 7, 2013
Cancer is a sine qua non for life as we know it.
There was a post today on National Geographic talking about a tumor that was found in a fossilized Neandertal's bone. It reminded me that I had written the piece below and hadn't found a home for it yet. The title, which is a big jarring, is:
Cancer is not a disease: there is no cure.
The way we describe things shapes and is shaped by the way that we think of them. Cancer has been described as a disease for as long as we have written record of medicine. It’s name comes from the greek word for crab, because the way it wedges itself into the host tissue is so like a crab wedges itself between rocks: inextricably.
The advent of the microscopic age at the turn of the 20th century brought with it an unprecedented view of the cellular level; anatomy and pathologies came into a new focus and a new science was born: microbiology. As physicians, we were offered a new opportunity to study diseases at the cellular level, to describe the panoply of new patterns that we saw under the microscope much like a team of explorers coming into undiscovered jungle filled with undescribed flora and fauna. We found that the cancers in each organ were not necessarily the same. Indeed, we found a rich diversity of cancers that could be reliably classified and whose prognoses and patterns of progression correlated. This richness of classification gave rise to the opportunity for disease specific treatment trials, and indeed accounted for most of our progress against these individual entities, and for the standard of care for most cancer types even today.
The dawn of the genomic age, first with the human genome project, and then with the cancer genome atlas, promised and delivered another wave of discovery and deeper, more detailed classification. Just as we can now tell how, and when two species of finch, or cave fish, diverged in their evolutionary history, so too can we tell when and how a tumor diverged from its tissue of origin. Early on in this story, we were tantalized by the discovery of specific genomic errors (mutations) that seemed to explain a cancer’s growth, and with the discovery of imatinib, a targeted ‘cure’ for a specific cancer (CML), and cancer seemed to be on its knees, ready to be cured.
The final cure, however, has continued to elude us. As we continue to discover more and more specific mutations, drug companies continue to develop specific drugs to target their action. Each of these drugs seems to work well in a subset of patients, for a time, but never provides the silver bullet that we have been promised, and ultimately fails in almost every case. The problem is that we are stuck in a paradigm where each disease has a cause, and each cause has a remedy. This linear thinking has dominated medicine, and indeed much of science, for most of human history, and has served us well. But continuing to think of cancer as a disease in this paradigm is not going to get us any closer to a cure - we have to shift our thinking and expectations, and embrace the reality that cancer is the result of a non-linear, highly degenerate process, and therefore has no 'cure'.
We have to shift our focus in the study of the cancer genome and stop trying to develop a comprehensive list of errors in the code that cause cancer, but instead learn the guiding principles behind the process - the equations of motion, if you will. Cancer is not a disease to be cured, but a pathologic condition of normal tissue evolving according to the very rules which allowed us to emerge from the primordial ooze. It is an inconvenient sine qua non for existence in our universe, as evolving, living organisms.
This reclassification is not intended to take anything away from those living with cancer, or who have suffered from it. Within any single patient, this pathologic condition has the capacity to cause as much, or more, suffering than does any other disease. And, as oncologists, our calling is to minimize this suffering, and when we can, cure our patient. But until we stop thinking about cancer as a disease that is the product of a linear process, and realize that it is a pathologic condition that is produced by any number of trajectories across an evolutionary landscape; until we stop looking for a single, silver bullet cure for all patients that doesn’t exist, we will continue to waste time that we could be using to understand the evolutionary dynamics of cancer so we can develop strategies to cure each patient.
Some further reading:
Exploiting ecological principles to better understand cancer progression and treatment
arXiv preprint
Basanta and Anderson
Cancer attractors: a systems view of tumors from a gene network dynamics and developmental perspective.
Huang et al. Semin Cell Dev Biol. 2009 Sep;20(7):869-76. doi: 10.1016/j.semcdb.2009.07.003
Oxidants, antioxidants and the current incurability of metastatic cancers.
Jim Watson, Open Biol. 2013 Jan 8;3(1):120144. doi: 10.1098/rsob.120144.
Wednesday, June 5, 2013
NIH Loan Repayment Program
When I was trying to decide what to do with my life after the Navy, it came down to two possibilities: medicine or physics. One, medicine, was wholly new to me but seemed like a good fit for my personality and it was a pretty sure bet, career wise. The other, physics, had been my passion since I was introduced to it in 1992 by Bob Shurtz, my high school AP Physics teacher (and the greatest physics teacher of all time). I had a really hard time picking between the two and ended up choosing medicine for pragmatic reasons, but always hoped to be able to incorporate my scientific inclincations with my new career. I found out pretty late in the game about the MD/PhD programs funded by the NIH through the Medical Scientist Training Program, but applied to a few anyways and was rejected. So, I started medical school as Case Western and was happy.
I distinctly remember one day during MS-1 when I was shadowing a hand surgeon in the OR at the Louis Stoke VA (our teaching VA in Cleveland), and the surgery resident, a PGY-2 in general surgery, described the NIH Loan Repayment plan to me. He was going to take a few years off after his PGY-3 year to pursue a PhD in tissue engineering. The benefit of doing this, he said, was that he could pick his clinical interest first, then pursue a related research area, rather than the MSTP folks who have to pick a scientific discipline before they know what sort of doctor they'll be. This has come into sharper focus for me as my best friend from medical school did his PhD in cardiac stem cells, assuming he would do cardiology, and fell in love with urologic surgery. It isn't so much that his training was wasted, it certainly wasn't, but he lost the momentum that he could have had had his clinical interests been aligned with his doctoral studies. So, in this sense, I'm glad that I wasn't accepted into the MSTP programs, as by going straight through medical school, I found my 'calling' as an oncologist, and then subsequently found how research fits into that - and this is an important distinction.
So, to continue the story, I chose radiation oncology, and now, between PGY-4 and 5, I have been taking time off to do dedicated translational research, and because of the NIH extramural loan repayment program, I am paying off a year of medical school for each year of qualified research! (Considering the opportunity cost of not practicing radiation oncology, I am still a bit 'behind' each year I postpone taking a faculty job, but this dulls the pain significantly.)
While I was disappointed that I didn't get in to the MSTP programs, I truly feel that, at least for me, this path is one that will better suit me for a rewarding career. Obviously, each person is different - had I had a burning passion about a certain kind of research early on, then following that path when I was younger might have been easier. I certainly face a whole slew of personal difficulties as a 37 year old PhD student with 2 young kids that I wouldn't have had I pursued this earlier. On the other side of the coin, with nearly ten years of medical experience, I bring a lot more to the table for my research than I would have after MS-2. At the end of the day, I am happy that the NIH funds both pathways, and we are in dire need of more physicians doing science. Like I said in my TEDMED talk, I truly feel that the physician-scientist plays a unique role in science. Indeed we are specifically trained to connect seemingly disparate pieces of data to build a theory, like a constellation of symptoms to build a diagnosis. As science gets more and more specialised, and knowledge silos get deeper and deeper, I feel this role will grow in importance as well - highlighting the need to keep these programs going.
While I love science and research, I came out of medical school with ~$200,000 in debt. I could not have afforded to take this time off if it weren't for this program. So, thanks NIH.
OK - so some nuts and bolts. You get a health related doctoral degree (specifics on eligibility here). Graduate. Get a job doing research at a non-profit, preferably NIH funded (post-doc sort of thing). They pay off your loans! Each quarter I get notified that they sent $8,750 off to Sallie Mae. The only continuing work, besides the research, is that your supervisor has to fill out a form saying you are working hard (thanks +Alexander Anderson) and you have to verify that the money is going where it is supposed to. Easy.
Here are some links that the folks at NIHLRP provided me to supply with data about who has gotten funded.
http://www.lrp.nih.gov/pdf/LRPFY2012DataBook.pdf
http://www.lrp.nih.gov/pdf/1211.1_trends_report.pdf
The application process was similar to any other grant writing endeavor. There was a 6 page project description as well as a biosketch and some institution-specific paperwork. Importantly, a major component is the training program that you and your mentor create, which must be described in detail in a letter. For me, this included a PhD training program, but this is not necessary. Here is my successful grant application:
You can then, competitively, renew this grant as well. I don't know how that process goes since I missed the renewal application cycle (whoops!). It turns out, they don't remind you... But, I'll be submitting a renewal application this year, as long as there isn't a 5 year gap, you can renew. This is important, as when I go back to finish my clinical training, I will not qualify, as I won't be able to dedicate at least 50% of my time to the research. However, once I am faculty again, as long as I have educational debt and am spending 50% of my time doing qualifying research, I can continue to apply.
Long winded, sorry. Short story: great program. Give it a shot! We need more health professionals doing research, and this is a great way to defray some of the opportunity cost of not practicing.
At press time - I realize I've forgotten a few key points. They are:
Application cycle runs September 1-Nov 15th. Funding level is nearly 50%, so your odds are good, and the initial contract period is 2 years. Go apply!
Also - here is a 'tip sheet' on best practices for applications: http://www.lrp.nih.gov/pdf/
I distinctly remember one day during MS-1 when I was shadowing a hand surgeon in the OR at the Louis Stoke VA (our teaching VA in Cleveland), and the surgery resident, a PGY-2 in general surgery, described the NIH Loan Repayment plan to me. He was going to take a few years off after his PGY-3 year to pursue a PhD in tissue engineering. The benefit of doing this, he said, was that he could pick his clinical interest first, then pursue a related research area, rather than the MSTP folks who have to pick a scientific discipline before they know what sort of doctor they'll be. This has come into sharper focus for me as my best friend from medical school did his PhD in cardiac stem cells, assuming he would do cardiology, and fell in love with urologic surgery. It isn't so much that his training was wasted, it certainly wasn't, but he lost the momentum that he could have had had his clinical interests been aligned with his doctoral studies. So, in this sense, I'm glad that I wasn't accepted into the MSTP programs, as by going straight through medical school, I found my 'calling' as an oncologist, and then subsequently found how research fits into that - and this is an important distinction.
So, to continue the story, I chose radiation oncology, and now, between PGY-4 and 5, I have been taking time off to do dedicated translational research, and because of the NIH extramural loan repayment program, I am paying off a year of medical school for each year of qualified research! (Considering the opportunity cost of not practicing radiation oncology, I am still a bit 'behind' each year I postpone taking a faculty job, but this dulls the pain significantly.)
While I was disappointed that I didn't get in to the MSTP programs, I truly feel that, at least for me, this path is one that will better suit me for a rewarding career. Obviously, each person is different - had I had a burning passion about a certain kind of research early on, then following that path when I was younger might have been easier. I certainly face a whole slew of personal difficulties as a 37 year old PhD student with 2 young kids that I wouldn't have had I pursued this earlier. On the other side of the coin, with nearly ten years of medical experience, I bring a lot more to the table for my research than I would have after MS-2. At the end of the day, I am happy that the NIH funds both pathways, and we are in dire need of more physicians doing science. Like I said in my TEDMED talk, I truly feel that the physician-scientist plays a unique role in science. Indeed we are specifically trained to connect seemingly disparate pieces of data to build a theory, like a constellation of symptoms to build a diagnosis. As science gets more and more specialised, and knowledge silos get deeper and deeper, I feel this role will grow in importance as well - highlighting the need to keep these programs going.
While I love science and research, I came out of medical school with ~$200,000 in debt. I could not have afforded to take this time off if it weren't for this program. So, thanks NIH.
OK - so some nuts and bolts. You get a health related doctoral degree (specifics on eligibility here). Graduate. Get a job doing research at a non-profit, preferably NIH funded (post-doc sort of thing). They pay off your loans! Each quarter I get notified that they sent $8,750 off to Sallie Mae. The only continuing work, besides the research, is that your supervisor has to fill out a form saying you are working hard (thanks +Alexander Anderson) and you have to verify that the money is going where it is supposed to. Easy.
Here are some links that the folks at NIHLRP provided me to supply with data about who has gotten funded.
http://www.lrp.nih.gov/pdf/LRPFY2012DataBook.pdf
http://www.lrp.nih.gov/pdf/1211.1_trends_report.pdf
The application process was similar to any other grant writing endeavor. There was a 6 page project description as well as a biosketch and some institution-specific paperwork. Importantly, a major component is the training program that you and your mentor create, which must be described in detail in a letter. For me, this included a PhD training program, but this is not necessary. Here is my successful grant application:
You can then, competitively, renew this grant as well. I don't know how that process goes since I missed the renewal application cycle (whoops!). It turns out, they don't remind you... But, I'll be submitting a renewal application this year, as long as there isn't a 5 year gap, you can renew. This is important, as when I go back to finish my clinical training, I will not qualify, as I won't be able to dedicate at least 50% of my time to the research. However, once I am faculty again, as long as I have educational debt and am spending 50% of my time doing qualifying research, I can continue to apply.
Long winded, sorry. Short story: great program. Give it a shot! We need more health professionals doing research, and this is a great way to defray some of the opportunity cost of not practicing.
At press time - I realize I've forgotten a few key points. They are:
Application cycle runs September 1-Nov 15th. Funding level is nearly 50%, so your odds are good, and the initial contract period is 2 years. Go apply!
Also - here is a 'tip sheet' on best practices for applications: http://www.lrp.nih.gov/pdf/
Subscribe to:
Posts (Atom)